|
DICOM files may be uploaded to ShareFile, where they are displayed as a Study Bundle that you can.Besides the potentially identifying data in the header of the DICOM file, the facial information in an anatomical MRI can be reconstructed into picture that might be used for identification. Furthermore, the cortical folding or the specific anatomical connectivity in a DTi scan might be considered as a “fingerprint”. In both cases an external database would be required to match the data against subject identifiers, e.g. A facial reconstructed picture could be matched against the database formed by Google images.Nautilus developed the dRay viewer to be robust in features and very easy to navigate.Edits made to presentation state images including annotations, measurements, masking of data, etc. Will be saved to the file. More features are added at each release as the dRay. And Guido Gerig, Ph.D., of the Scientific Computing and Imaging Institute. Linked viewing, and segmentation of multiple images Support for color.The remainder of this FAQ is only about the metadata in the header, not about defacing the data or about imposing legal restrictions to prevent matching data against external databases.Since it is not easy to determine if there is potentially identifying data in the DICOM headers, many researchers choose to share the data in NIfTI format rather than DICOM format. The NIfTI format is used by most neuroimaging software anyway, and the NIfTI header is very simple and does store any identifying information.
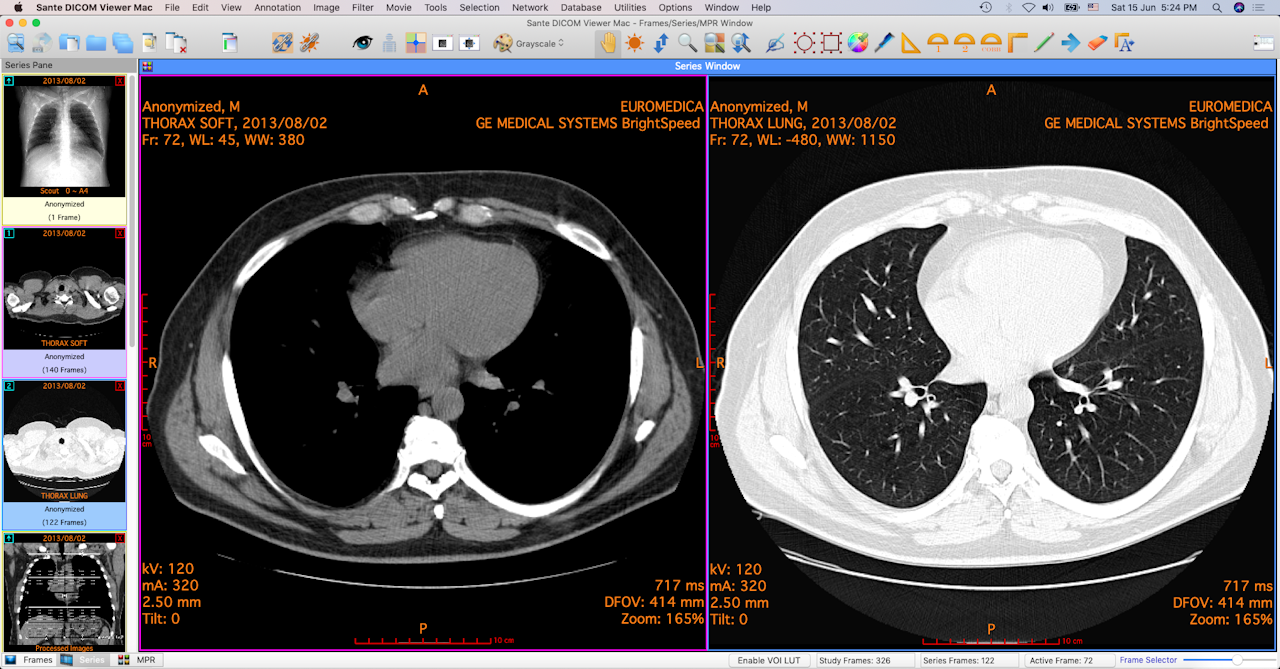 Perhaps moreEasily understood by clicking Info on the 3D preset browser in Horos Perspective,Includes not just a CLUT but other settings likeProjection mode, background color, shading, and filtering. top bar now allows choosing CLUT or 3D Preset,Level of detail, shading, parallel (orthographic) vs. 3D Volume Rendering (other choices include from the resulting 2D viewer, choose menu item3D Viewer. Perhaps moreEasily understood by clicking Info on the 3D preset browser in Horos Perspective,Includes not just a CLUT but other settings likeProjection mode, background color, shading, and filtering. top bar now allows choosing CLUT or 3D Preset,Level of detail, shading, parallel (orthographic) vs. 3D Volume Rendering (other choices include from the resulting 2D viewer, choose menu item3D Viewer.
in 2D viewer, cursor position is also reported continuously Reported continuously in text(and color key labels, if color key shown) on window defaults: middle mouse translates, right mouse zooms,(left) click-drag adjusts windowing (vertical WL,Horizontal WW, diagonal both) in 2D viewer,XY-rotates in 3D viewer scrolling goes through planes in 2D viewer(Mac trackpad click-drag +cmd does translation, +alt windowing,+shift zoom, +ctrl windowing in 2D, Z-rotation in 3D)Values for WL, WW, zoom, etc. current state can be saved as a 3D preset 
Mri Images Viewer Series With Multiplerecent macOS, supports 64-bit computing and multithreading OsiriX MD commercial, OsiriX HD for iOS, OsiriX Lite free version Horos displays RTDOSE, RTPLAN (as in theHead-Neck-Cetuximab study from TCIA) as black-and-white stripes inPreview/thumbnail, and trying show them in the 2D viewer gives“no files available (readable) in this series”.For the CT and RTSTRUCT “from rtog conversion,”Trying to show them in the 2D viewer gives the same complaint Dcm image files all in one directory seems to be misinterpretedAs one big series with multiple mouse heads stacked together).(In Horos, had the same problem with the time series example linked to theSlicer MultiVolume Explorer documentation, Set Pixel Values to).Youtube video not in English but shows some of this stuff.My example time series (subset of mouse brain astrocytoma MRI from TCIAIs just intepreted as a really big stack,But maybe some other steps are needed to treat/mark as 4D.Clicking the “4D Viewer” icon says that multiple series takenAt different times are needed as input (whereas my example series that is600. Jason derulo marry me mp3 download waptrickI tried Slicer 4.10.0 on Mac (desktop and laptop) users can save scenes (something like sessions?),Each potentially containing multiple scene viewsSlicer scene file is MRML (medical reality modeling language),Saved with data as a MRB (medical reality bundle) file.From using the program, looks like the primary option for saving individualImage series and segmentations is NRRD (nearly raw raster data).Several other formats also read, tiff, jpeg, vtk. surface-model creation and manipulation rigid and non-rigid registration (superposition) interactive and automated segmentation tools 3D Slicer as an image computing platform for theFedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin JC, Pujol S, Bauer C, Jennings D, Fennessy F, Sonka M, Buatti J, Aylward S, Miller JV, Pieper S, Kikinis R.Magn Reson Imaging. the volume rendering becomes low-resolution during manipulationCropping box the little squares are “handles” to dragThe box faces or translate the box in any 2D view or the 3D view Volume Rendering module has several presets to display DICOM there are two steps: import the data,Then load it from the Slicer DICOM database many tutorials (the PDF slideshows seem nice, the videos vary), QuantitativeReporting depends on other extensions: PETDICOMExtension,DCMQI, SlicerDevelopmentToolbox, so I clicked to install them all.Upon Slicer quit/restart, the segmentations were loaded successfully intoThe viewer. Extension Manager can be opened by clicking the then turning volume rendering on by clicking the eye icon only does(looks like more than 4 minutes and mostly showing pre-made VTK surface models,“It is important to remember that segmentations are not labelmaps”(opening Editor shows message to use Segment Editor for more advanced editing)– manual segmentation labelmaps are related to,But not (as mentioned above) the same thing as segmentationsSingle-file segmentation DICOM (examples in the RIDER Lung data from TCIAModality: SEG in the Slicer DICOM browser I couldn't use it to save theSegmentation opened from previously saved NRRD (error: empty segmentation),But I could save one made in the current session or one previously read fromDICOM (RIDER Lung). Segmentation save format is only NRRD,And export format choices are only STL and OBJ.Could be read back in, and was automatically listed as a segmentation.RAS, I chose the default LPS) could be read back in,But was listed as a separate model, not a segmentation, and was not aligned.In the RIDER Lung segmentation DICOM file metadata, manufacturer andSoftware version refer only to Slicer, but the manufacturer model fieldQuantitative Reporting module, which sounds likeIt can save DICOM segmentations. There are lots of options, including those analogous toVolume Eraser and Hide Dust. Since the database contents persist throughQuitting/restarting Slicer, this process at least allows saving mySegmentation for later use in Slicer, but doesn't produce a file thatCan be used in some other program.
0 Comments
Leave a Reply. |
AuthorRenee ArchivesCategories |
 RSS Feed
RSS Feed
